Computable knowledge Part 2 of 2: Learning from the HDR UK collaborathon – make it FAIR!
26 February 2021 | Author: Philip Scott, Researcher
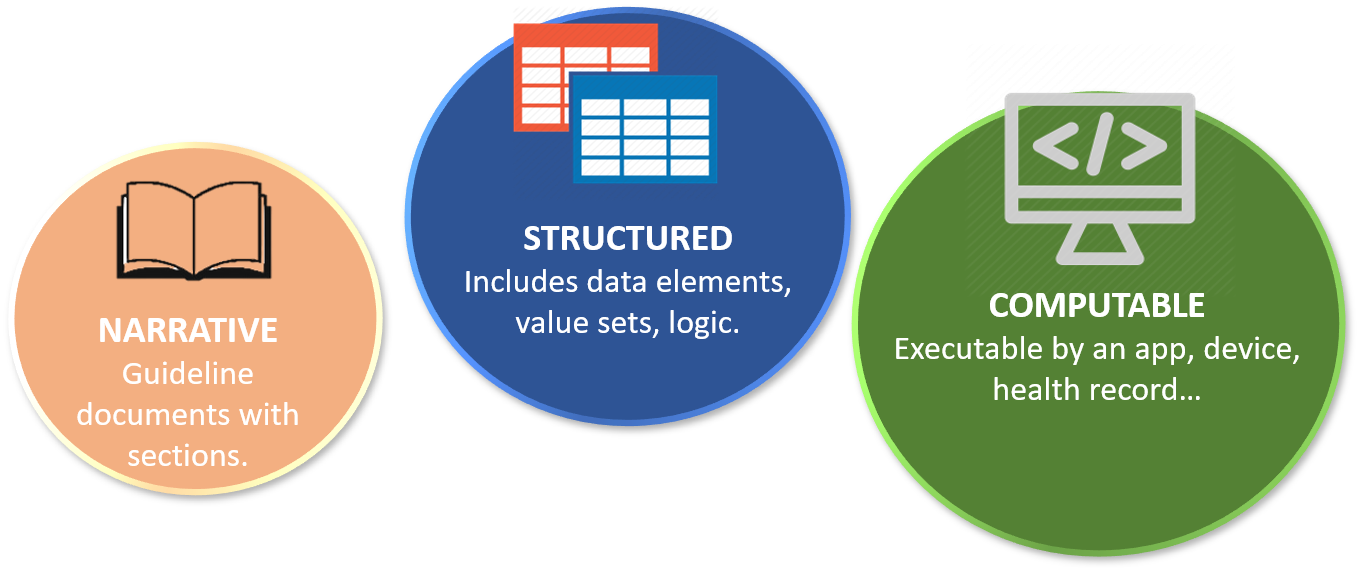
The previous blog article explained the concept of computable knowledge and why we need it in Learning Health Systems (LHS). Now we report on a recent HDR UK Better Care Catalyst initiative to promote the concept of a knowledge ecosystem and explore some of the practical issues to achieving it.
HDR UK has already recognised the potential of the LHS model, and funded the Better Care National Priority which supports LHS development and implementation in a network of healthcare organizations across the country. In partnership with The Health Foundation, there are also three Better Care Catalyst workstreams to support the creation of an enabling environment which maximises the opportunity to embed this approach more widely.
In December 2020, we held the first ever computable knowledge “collaborathon”, as a virtual online event. The event aimed to raise awareness of standard ways to exchange knowledge between different computer applications and to provide a learning opportunity for clinical, academic and industry participants to use computable knowledge. In the planning stage, we explored various possibilities of computable knowledge representation based upon who was willing to take part, but finally selected FHIR (Fast Healthcare Interoperability Resources, http://hl7.org/fhir/) as it is globally popular and supports both data and knowledge artefacts, and CQL (Clinical Quality Language, https://cql.hl7.org/) as it is being co-developed with FHIR and promoted in US policy (which has a powerful effect on the healthcare IT market).
We published an open call for participation and recruited four teams made up of a total of 11 participants (4 female, 7 male) from 7 different institutions: UCLH, Imperial College, NHS Lothian, University of Portsmouth, University of Manchester, Ulster University and University Hospitals Geneva. Teams had a choice of (1) developing a knowledge artefact in CQL/FHIR based on a selection of NICE guidelines or (2) implementing existing knowledge objects from international repositories. We had an outstanding judging panel, comprising senior clinicians, chief clinical information officers, and a director from NICE.
All the teams chose to work on producing CQL artefacts from existing guidelines, with feedback suggesting that creating your own knowledge object is perceived as easier than re-using someone else’s. Three teams worked on acute kidney injury detection , and one group worked on COVID-19 diagnosis. To do this, they had to identify the implicit algorithm in their source guideline, work out the key data items needed to execute the algorithm and identify the particular SNOMED CT values to trigger the decision points. For example, here is a simplified CQL fragment from one team’s work:
codesystem “SNOMED”: ‘http://snomed.info/sct’code “Cough (SNOMED)”: ‘49727002’ from “SNOMED” display ‘cough SNOMED’define “HasCough”:exists(ActiveCondition([Condition: “Cough (SNOMED)”]))
Translated, the first line means that “SNOMED” is used as shorthand for the SNOMED CT terminology. The next line creates the label “Cough (SNOMED)” to mean the SNOMED CT concept ID 49727002. The last two lines identify when a patient “HasCough”, which is triggered if the patient record has the appropriate SNOMED code for cough. In itself this is trivial, but the CQL logic combined with the FHIR data structures allows construction of complex decision rules. Even with a basic knowledge of a programming language teams were able to produce workable prototypes.
What did we learn from the event about computable knowledge? NICE supports the vision of computable knowledge guidelines, and although narrative content is still predominant right now, they can see the need for different ways of working in the future. HDR UK strongly promotes the Better Care paradigm, using data and analytics to inform care, and the re-use of actionable insights to enable scaling.
Despite being seen as a priority, there are few examples of computable knowledge artefacts reported across the Better Care and wider research community that are applicable to frontline clinical decision-making. Many projects publish their software code in open repositories such as github, but that is (with one notable exception) typically for data processing not knowledge implementation. There is excellent work on phenotype definitions, which along with other curated code lists and ontologies are foundational to computable knowledge. Some researchers are developing apps to implement the knowledge they produce, but there is apparently limited awareness of the concept of sharing computable knowledge beyond the original project.
Why is turning research into computable knowledge so challenging? Fundamentally, the research focus seems still to be largely about traditional publication and that is the main basis for assessment and incentives.
What does HDR UK need to do, at the organisational level, to promote the production of re-usable computable knowledge? Firstly, establish a cross-cutting knowledge management work group that will leverage the domain-specific expertise from the Hubs and more generalist knowledge engineers with public involvement support. They can co-design a knowledge management strategy and a list of specific and general recommendations such as aiming to extend the principles for participation (https://www.hdruk.ac.uk/wp-content/uploads/2020/03/200304-Principles-for-Participationv2pdf.pdf) so that not only data but also knowledge must be FAIR: findable, accessible, interoperable and re-usable. Secondly, project proposals should be required to specify what open standards will be employed to achieve the FAIR principles. Thirdly, projects should be required to contribute at least one computable artefact into an annual collaborathon. Fourthly, we should work with real-world clinical systems with active supplier co-design. Finally, we must ensure public involvement in the development of a computable knowledge strategy – there are important issues of trust, governance and curation that demand the direct voice of citizens and a commitment to an open and collaborative knowledge management strategy.